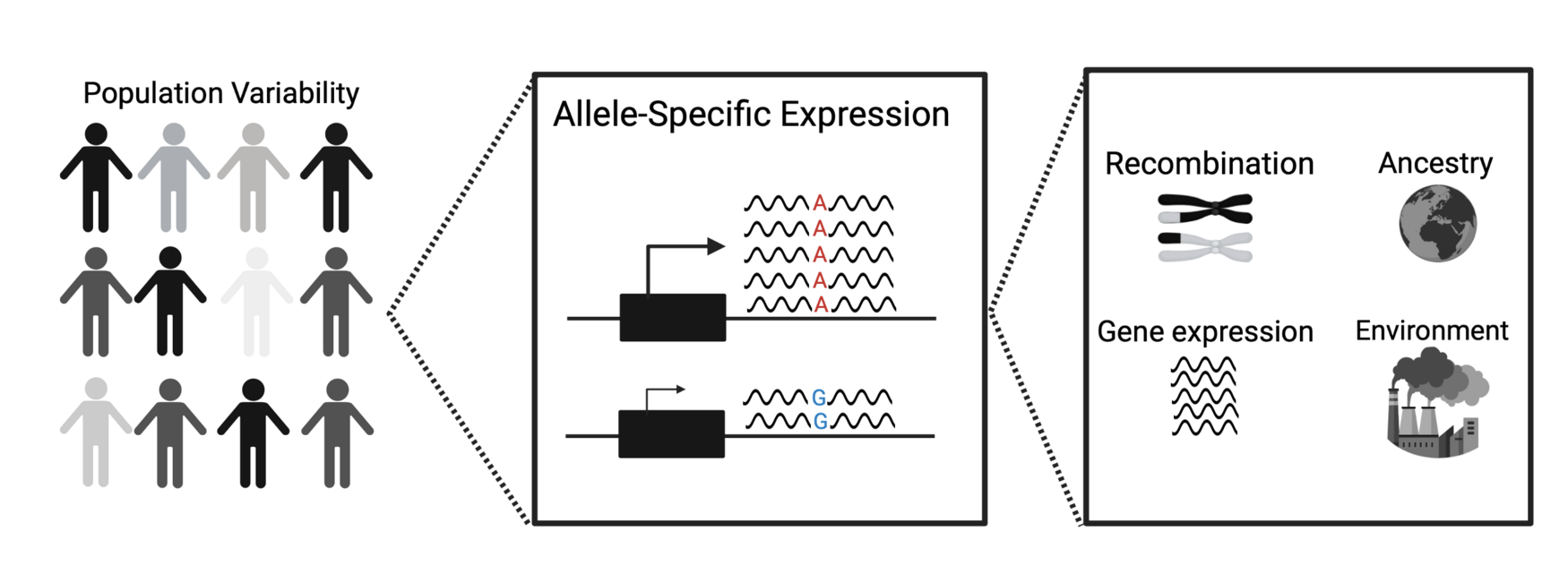
A person’s physical traits and susceptibility to disease tend to largely depend on gene expression and its regulation. Gene expression is the process of how the cell decodes a gene to produce proteins. Changes in this process can majorly impact human health, both on an individual and population level, as it alters proteins that the human body depends on to operate normally. One prominent type of gene expression is allele-specific expression (ASE), which occurs in 5-15% of the coding region of the genome. Researchers define ASE as an imbalanced or unequal expression leading to different transcript levels between the paternal and maternal alleles or copies of a gene.
In their latest publication in Science Advances, MoGen Ph.D. student Michelle Harwood and other members of Dr. Philip Awadalla’s lab at OICR demonstrate how genetic recombination and other factors influence the ASE of deleterious and disease-associated mutations in human populations. Deleterious mutations either produce no or dysfunctional proteins, potentially resulting in disease or increased risk of it. The team specifically analyzed the genomes of three French-Canadian sub-populations, one general African and one general European population, to investigate ASE genome-wide.
Proteins create or influence the phenotype, or external traits, where evolutionary forces such as natural selection then act on them and influence how frequent specific alleles and mutations are in a population. The team confirmed that ASE reduces the expression of deleterious mutations and suggested the strength of natural selection is the basis of this. They then unearthed that ASE occurs most frequently in genomic regions with increased recombination rates and gene expression levels or the number of transcripts. Recombination is a process that increases the number of haplotypes, or groups of linked alleles that get inherited together, and a population’s genetic diversity by breaking apart DNA and recombining them to make new allele combinations. Harwood summarizes the process as natural selection acting more efficiently in regions with higher recombination and expression levels, resulting in more ASE occurring and minimizing the expression of deleterious or disease-associated mutations. A related but similarly critical finding by the team revealed that historically older populations, such as Europeans and Africans, regulate ASE and disease harbouring alleles better relative to younger populations like French Canadians.
The group also found that environmental factors regulate and influence ASE. The team determined this by contrasting allele expression levels for French-Canadian individuals that have moved or changed residencies relative to their ancestors with whom they share common genetic backgrounds. They found ASE in specific genes differs between people having the same ancestry but who lived in different geographic environments. As described in a previous study from the Awadalla lab, environmental influences known to alter gene expression, such as pollution levels, could play a role.
The results also sparked interesting questions surrounding ASE in immune-related genes. One interesting finding was the variable ASE in the GBP3 gene, which has antiviral activity against influenza. “The variable ASE may not be from ancestry or [living] environment but rather the immune response,” Harwood notes when discussing this finding, “perhaps some individuals were recovering from or undergoing an infection at the time of sampling, which could explain variable ASE in these immune genes.” The role of ASE in immunity is Harwood’s current research direction, specifically to analyze ASE on a single-cell level in blood.
In conclusion, ASE under-expresses deleterious and disease-associated mutations most efficiently in areas of the genome where selection is most efficient, namely regions with high recombination rates and expression levels. These findings have important implications from multiple perspectives in evolutionary and human genetics. It also demonstrates the potential importance of external factors such as the living environment and the immune response in regulating gene expression. Knowing how the genome and environment regulate these variants and how and why some individuals carry a mutation but don't exhibit disease is vital in potentially treating and preventing these conditions.
A big thank you to Michelle and Dr. Awadalla for providing their expertise and insight!
Read the article here: Harwood MP, Alves I, Edgington H, Agbessi M, Bruat V, Soave D, Lamaze FC, Favé M-J, Awadalla P. Recombination affects allele-specific expression of deleterious variants in human populations. Science Advances. 2022;8(19):eabl3819. doi:10.1126/sciadv.abl3819